Really, you had me fooled! Where did you learn to use bioinformatic tools? I'll be impressed if that is self-taught in anyway.
Yup, self-taught.
As Armando posted above, it's a pretty well studied that CAG repeat length and sensitivity of the androgen receptor are inversely correlated. That is people with small CAG repeats will have a more sensitive androgen receptors and those with longer CAG repeats will have an "insensitive" androgen receptor (
https://www.ncbi.nlm.nih.gov/pubmed/12641825). Although this has been shown in many studies to affect things like bone density, reproductive functions, and cardiovascular risks, the role of CAG repeats and sensitivity in male pattern baldness has a bit of conflicting evidence.
This is probably because hair loss is very polygenic (many genes involved) and the sensitivity of the AR may only play a role in some peoples hair loss.
Very polygenic indeed. Have you seen the recent GWAS by
Hagenaars et al.? With a sample of 52,000 men, they found 287 variants around ~100 genetic regions. I combined their results with those from
this meta-analysis and from
this study on the genetics of 42 traits. Based on (1) proximity of gene to SNP, (2) known role in hair biology, (3) known interactions with other associated genes, and (4) some other tricks for specific cases, I picked the most plausible gene from each region and came up with this:
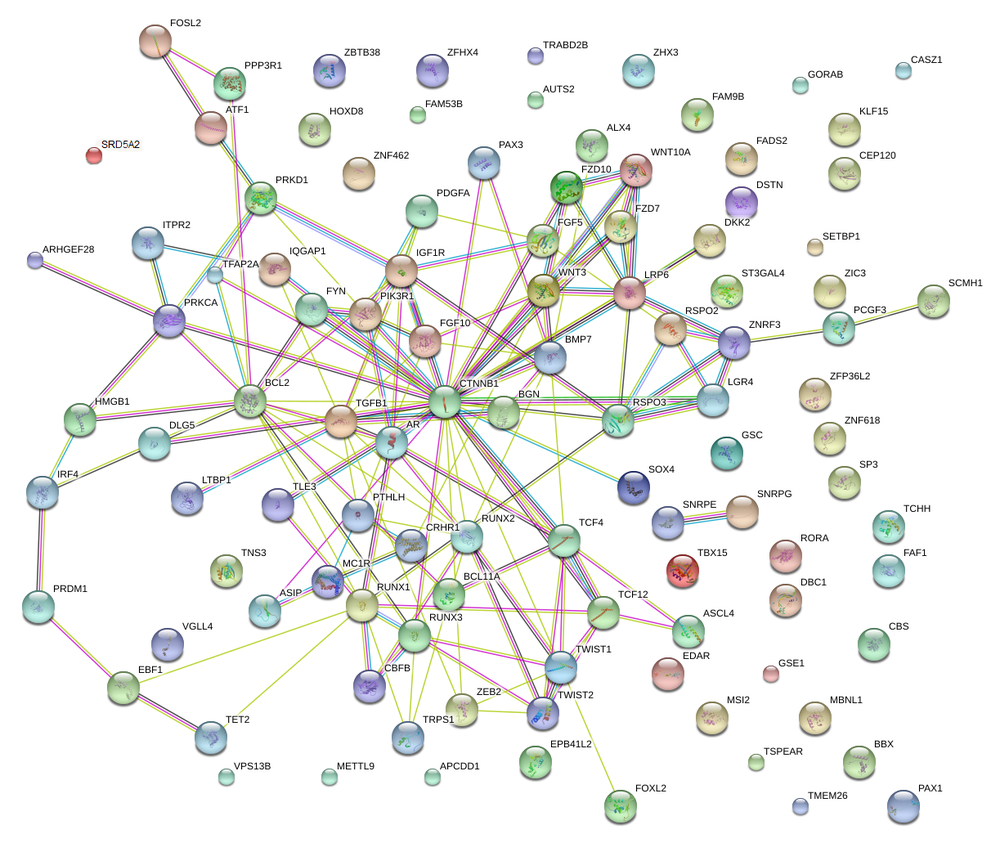
So the best theory on why this may work on those with shorter CAG repeats may be by the Andrologist Michael Zitzman. He says the following “Men with shorter CAG repeat tracts respond significantly better to this treatment than those with longer CAG repeats. However this may not be pharmacogenetics but rather a selection phenomenon: the men with shorter repeat tracts are probably those with androgenically induced baldness and this are more susceptible to a reduction of DHT”
I tend to believe this theory. Some young men will be more susceptible to hair loss because they have “sensitive” hair loss follicles in part due to this CAG repeat region. So when we take away the main cause of hair loss for them, DHT binding to the AR, it returns them to a state where hair is freer to grow normally. Men who have hair loss and longer CAG repeats may have essentially a different factor playing a larger role in their hair loss, such as in the WNT signalling pathway.
I was thinking something along those lines, but given that we know androgens are required for M.P.B to progress, what I thought of was a little different. Consider this:
The 42 traits study found 50 genetic regions associated with the "unibrow" trait. 17 of these* are shared with A.G.A. The vast majority of these don't tag the same variant (or variants in high LD with the A.G.A variants). I assume this corresponds to the same genes being under the control of different enhancers depending on position in the body? In any case, to me this suggests that mechanisms involved in A.G.A are similar to mechanisms that determine "normal" hair growth.
* These are the regions near TBX15, EDAR, the HOXD cluster, PAX3, FGF5, IRF4, RUNX2, PRDM1, RSPO3, TWIST1, AUTS2, ZFHX4, ALX4, MC1R, PAX1, BMP7, and CRHR1.
Let's say for the sake of argument that AR can cause HF miniaturization through something like the following:
AR -> affects activities of dermal papilla transcription factors like Twist, Runx, Trps1, etc. -> inhibits secreted factors like R-spondins -> decreased Wnt pathway activation in HF stem cells -> decreased activation of HF stem cells -> HF miniaturization
Several of the SNPs around these DP transcription factors disrupt E-box motifs (CANNTG, which basic helix-loop-helix TFs like Twist1, Twist2, Tcf4, and Tcf12 bind to) including around Twist2 and Tcf4 themselves. A couple SNPs near RSPO2 -- one protective and one harmful -- disrupt E-box motifs. Therefore, there could be self-contained regulatory loops among these transcription factors affecting secreted factors like R-spondins. And let's say that past a certain threshold of activitation AR permanently rewires this network to reduce hair growth. This would depend both on the level of AR activation (which would depend on genetics) and the "switching" threshold for these downstream factors (which would also depend on genetics). Then you could maybe have acute effects on AR through this network, and also permanent effects through it's own internal regulatory loops, the latter of which would be analogous to the regulation of "normal" hair growth. If that were the case, we might find the following groups of A.G.A sufferers:
1) Those with a high level of AR activity, and low switching threshold among the downstream factors. Result: Aggressive hair loss poorly reversed by anti-androgens.
2) Those with a high level of AR activity, and moderate to high switching threshold among the downstream factors: Result: Anywhere from mild to aggressive hair loss. Better reversal of recent hair loss with anti-androgens.
3) Those with a moderate level of AR activity, but low switching threshold among the downstream factors. Result: Androgens still required to progress. Anywhere from mild to aggressive hair loss poorly reversed by anti-androgens.
(The opposite pattern might hold true for androgen-dependent development of facial hair and body hair)
Comparing the CAG repeats would select against the third group, and may also select against those with SNPs near AR that would presumably increase AR expression.
Is that at all plausible or does it sound like total bullshit to you?

It is clear much research needs to be done in this area. I’d love to do it and the more I think about it, it seems like funding is going to come from this community rather than research grants. Scientists just don’t take hair loss seriously enough.
Yes, the amount of research going into A.G.A is a joke. Good luck to you if you do pursue research into this.